Building a RAG Application with pgvector and the OpenAI API
Retrieval-Augmented Generation is now the default architecture for adding knowledge to LLM-powered applications. The idea is simple: when a user asks a question, retrieve relevant context from your own data, then pass that context to the model alongside the question. The model answers based on what you gave it, not just what it was trained on.
Most RAG tutorials reach for LangChain or LlamaIndex immediately. Those are fine frameworks. But they add abstractions between you and what is actually happening, and when something breaks in production, you end up debugging framework internals instead of your own code.
This guide builds a complete RAG pipeline from scratch using three things: pgvector for vector storage and retrieval, the OpenAI Python SDK for embeddings and generation, and psycopg for the database connection. No orchestration layer. Every line does exactly what it looks like.
If you want the LangChain version, see our pgvector + LangChain guide. This post is for the cases where you want direct control.
What We Are Building
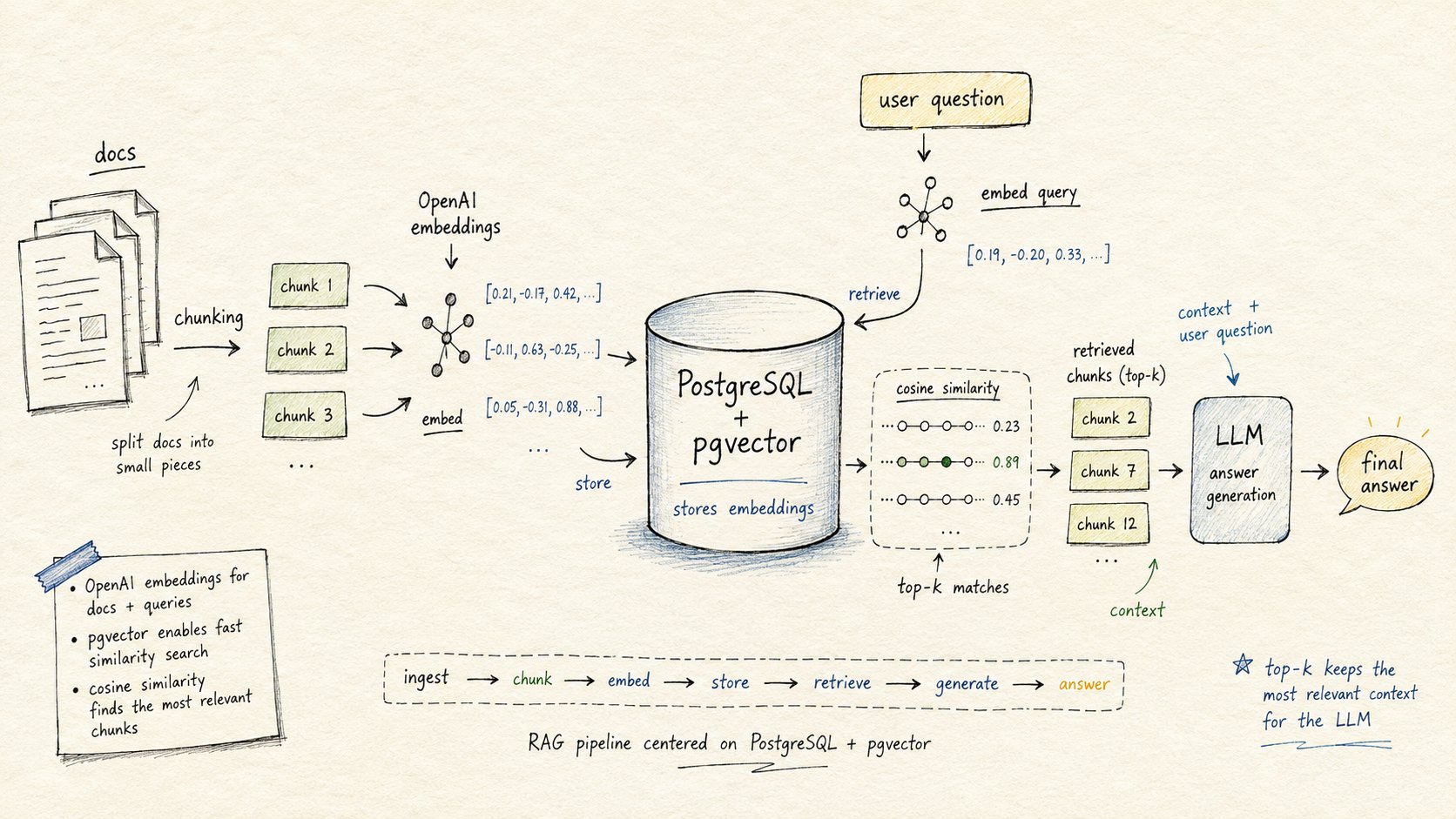
A document Q&A system. You load documents (plain text, PDF extracts, whatever), split them into chunks, embed each chunk with OpenAI, store the embeddings in pgvector, then answer questions by:
- Embedding the question
- Finding the most semantically similar chunks via vector search
- Passing those chunks to a chat model as context
- Returning the generated answer
The whole thing fits in about 150 lines of Python. Let's go through each part.
Prerequisites
You need:
- PostgreSQL with the
vectorextension (any Rivestack database has it enabled by default) - Python 3.10+
- An OpenAI API key
Install the dependencies:
pip install openai psycopg psycopg[binary] python-dotenvWe use psycopg (the v3 driver) rather than psycopg2. The async support and binary protocol are worth the switch for new projects.
Set your environment variables:
export OPENAI_API_KEY="sk-..."
export DATABASE_URL="postgresql://user:password@host:5432/dbname"Setting Up the Database
Enable the extension and create the tables:
CREATE EXTENSION IF NOT EXISTS vector;
CREATE TABLE documents (
id BIGSERIAL PRIMARY KEY,
title TEXT NOT NULL,
source TEXT,
created_at TIMESTAMPTZ DEFAULT now()
);
CREATE TABLE chunks (
id BIGSERIAL PRIMARY KEY,
document_id BIGINT REFERENCES documents(id) ON DELETE CASCADE,
content TEXT NOT NULL,
token_count INTEGER,
embedding VECTOR(1536),
created_at TIMESTAMPTZ DEFAULT now()
);The separation between documents and chunks is important. One document produces many chunks. You want to be able to delete a document and have its chunks disappear automatically via the cascade. You also want to store metadata at the document level (title, source URL, date) without repeating it on every chunk row.
The VECTOR(1536) dimension matches text-embedding-3-small. If you switch to text-embedding-3-large, you would use VECTOR(3072).
After inserting data, create the HNSW index for fast approximate nearest neighbor search:
CREATE INDEX ON chunks USING hnsw (embedding vector_cosine_ops)
WITH (m = 16, ef_construction = 64);Do not create the index before you have data. The HNSW build reads the entire table. Creating it on an empty table and then inserting rows works but gives you no benefit during insertion and forces you to wait for the index to build up incrementally. See our HNSW tuning guide for the full parameter breakdown.
Chunking Documents
How you split documents matters more than people expect. Chunks that are too small lose context. Chunks that are too large dilute the embedding signal (the model averages over too many concepts) and blow up your context window during generation.
A practical starting point: 400 tokens per chunk, 50-token overlap between adjacent chunks. The overlap prevents you from splitting a sentence at a boundary and losing the coherent meaning of the idea that spans the split.
import re
from typing import Iterator
def chunk_text(text: str, chunk_size: int = 400, overlap: int = 50) -> Iterator[str]:
words = text.split()
start = 0
while start < len(words):
end = min(start + chunk_size, len(words))
yield " ".join(words[start:end])
if end == len(words):
break
start += chunk_size - overlapThis is a simple word-based splitter. For production, you would use a token-aware splitter (the tiktoken library counts tokens the same way the OpenAI API does) and handle paragraph and sentence boundaries. But for most documents, word-based chunking at 400 words gets you close enough to start.
Generating and Storing Embeddings
Now the core of the ingestion pipeline:
import os
import psycopg
from openai import OpenAI
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
db_url = os.environ["DATABASE_URL"]
EMBEDDING_MODEL = "text-embedding-3-small"
EMBEDDING_DIMENSIONS = 1536
def embed_texts(texts: list[str]) -> list[list[float]]:
response = client.embeddings.create(
input=texts,
model=EMBEDDING_MODEL,
)
return [item.embedding for item in response.data]
def ingest_document(title: str, source: str, text: str) -> None:
chunks = list(chunk_text(text))
# Embed all chunks in one API call (max 2048 inputs per call)
batch_size = 256
all_embeddings = []
for i in range(0, len(chunks), batch_size):
batch = chunks[i : i + batch_size]
all_embeddings.extend(embed_texts(batch))
with psycopg.connect(db_url) as conn:
with conn.cursor() as cur:
cur.execute(
"INSERT INTO documents (title, source) VALUES (%s, %s) RETURNING id",
(title, source),
)
doc_id = cur.fetchone()[0]
for content, embedding in zip(chunks, all_embeddings):
cur.execute(
"""
INSERT INTO chunks (document_id, content, token_count, embedding)
VALUES (%s, %s, %s, %s)
""",
(doc_id, content, len(content.split()), embedding),
)
conn.commit()
print(f"Ingested '{title}': {len(chunks)} chunks")A few things worth noting here.
Batching the embedding calls is important. Calling the API once per chunk is slow and wastes request overhead. The embeddings.create endpoint accepts up to 2048 inputs per call. Batching 256 at a time is a safe middle ground that keeps latency low without hitting limits.
The psycopg driver handles the conversion of Python lists to the vector type automatically when the column type is known. No manual serialization needed.
Retrieval: Finding Relevant Chunks
When a question comes in, embed it and run a cosine similarity search:
def retrieve_chunks(question: str, limit: int = 5) -> list[dict]:
question_embedding = embed_texts([question])[0]
with psycopg.connect(db_url) as conn:
with conn.cursor() as cur:
cur.execute(
"""
SELECT
c.content,
d.title,
d.source,
1 - (c.embedding <=> %s::vector) AS similarity
FROM chunks c
JOIN documents d ON d.id = c.document_id
ORDER BY c.embedding <=> %s::vector
LIMIT %s
""",
(question_embedding, question_embedding, limit),
)
rows = cur.fetchall()
return [
{"content": row[0], "title": row[1], "source": row[2], "similarity": row[3]}
for row in rows
]The <=> operator computes cosine distance. Ordering by distance ascending gives you the most similar chunks first. The 1 - distance conversion turns it into a similarity score between 0 and 1, which is easier to reason about when you want to filter out low-quality matches.
You can optionally add a similarity threshold to discard chunks that are not actually relevant:
WHERE 1 - (c.embedding <=> %s::vector) > 0.75A threshold of 0.7 to 0.8 works for most use cases. Below 0.7, you are often retrieving chunks that are tangentially related at best. Tune this against your own data.
Filtering by Metadata
One of the real advantages of pgvector over dedicated vector stores is that you can combine vector search with any SQL condition. Want to restrict retrieval to a specific document, a date range, or a user's private documents?
SELECT c.content, d.title, 1 - (c.embedding <=> %s::vector) AS similarity
FROM chunks c
JOIN documents d ON d.id = c.document_id
WHERE d.source = 'internal-wiki'
ORDER BY c.embedding <=> %s::vector
LIMIT 5No special metadata indexing, no separate filter API, no eventual consistency lag. It is just a WHERE clause.
Generation: Answering the Question
Take the retrieved chunks, format them as context, and call the chat completion API:
CHAT_MODEL = "gpt-4o-mini"
def answer_question(question: str, limit: int = 5) -> str:
chunks = retrieve_chunks(question, limit=limit)
if not chunks:
return "I could not find relevant information to answer that question."
context = "\n\n---\n\n".join(
f"Source: {c['title']}\n{c['content']}" for c in chunks
)
system_prompt = (
"You are a helpful assistant. Answer the user's question using only the "
"context provided below. If the answer is not in the context, say so clearly. "
"Do not make up information.\n\n"
f"Context:\n{context}"
)
response = client.chat.completions.create(
model=CHAT_MODEL,
messages=[
{"role": "system", "content": system_prompt},
{"role": "user", "content": question},
],
temperature=0.2,
)
return response.choices[0].message.contentA few decisions made here:
Low temperature. RAG answers should be grounded in the retrieved context. You want factual accuracy, not creativity. A temperature of 0.1 to 0.3 keeps the model close to the source material.
Explicit grounding instruction. Telling the model to answer only from the provided context and to say when it does not know significantly reduces hallucination. Without this, models will sometimes blend retrieved context with their training data in ways that are hard to detect.
Context formatting. Separating chunks with --- dividers and labeling each with a source makes it easier for the model to reason about which chunk is answering which part of the question. This also makes it easier to implement citations later.
Putting It All Together
if __name__ == "__main__":
# Ingest a sample document
sample_text = """
PostgreSQL is an open-source relational database system. It supports advanced
data types and has strong ACID compliance. pgvector extends PostgreSQL with
vector similarity search, enabling AI applications to store embeddings alongside
relational data. This eliminates the need for a separate vector database in most
production scenarios. The extension supports both exact and approximate nearest
neighbor search via IVFFlat and HNSW indexes.
"""
ingest_document(
title="PostgreSQL and pgvector Overview",
source="internal-docs",
text=sample_text,
)
# Answer a question
question = "What are the index types supported by pgvector?"
answer = answer_question(question)
print(f"Q: {question}\nA: {answer}")Run it and you get a complete round-trip from document to answer.
Production Considerations
Batch Ingestion Performance
For large document collections, run ingestion concurrently. The OpenAI embeddings API supports parallelism and has generous rate limits. Use asyncio with psycopg's async connection to pipeline the database writes:
import asyncio
import psycopg
async def ingest_chunks_async(conn, doc_id: int, chunks: list[str], embeddings: list[list[float]]):
async with conn.cursor() as cur:
await cur.executemany(
"INSERT INTO chunks (document_id, content, embedding) VALUES (%s, %s, %s)",
[(doc_id, chunk, emb) for chunk, emb in zip(chunks, embeddings)],
)
await conn.commit()Re-embedding After Model Upgrades
When OpenAI releases a new embedding model (or you switch from text-embedding-3-small to text-embedding-3-large), your existing embeddings are no longer comparable. You need to re-embed everything.
Track the embedding model and version on each row:
ALTER TABLE chunks ADD COLUMN embedding_model TEXT DEFAULT 'text-embedding-3-small';Then you can identify and re-process stale rows without dropping the whole table:
SELECT id, content FROM chunks WHERE embedding_model != 'text-embedding-3-large';Retrieval Quality
The default retrieval (top 5 chunks by cosine similarity) works for simple question-answer tasks. For complex questions that span multiple topics, consider:
Query rewriting. Ask the model to rephrase the user's question before embedding it. This often produces better embeddings for retrieval because it removes ambiguity and expands abbreviations.
Reciprocal rank fusion. Run multiple queries (original, rewritten, keyword variants) and merge the result lists using RRF before sending context to the model. This is the same approach described in our hybrid search guide.
Re-ranking. Retrieve a larger candidate set (top 20) and re-rank with a cross-encoder before trimming to the final context window. Cross-encoders are slower than embedding similarity but more accurate.
Context Window Management
Every model has a context limit. For gpt-4o-mini that is 128,000 tokens, which is generous. But if you pass too many chunks, you dilute the signal. More context does not always mean better answers. Studies suggest models perform best with 3 to 8 highly relevant chunks, not 20 loosely relevant ones.
Track token counts on your chunks (the ingestion code above stores token_count) and limit by tokens rather than chunk count:
def build_context(chunks: list[dict], max_tokens: int = 3000) -> str:
parts = []
used = 0
for chunk in chunks:
tokens = chunk.get("token_count", len(chunk["content"].split()))
if used + tokens > max_tokens:
break
parts.append(f"Source: {chunk['title']}\n{chunk['content']}")
used += tokens
return "\n\n---\n\n".join(parts)Running on Managed PostgreSQL
Self-hosting PostgreSQL and keeping the pgvector extension up to date is operational work that gets more complex as you scale. Connection limits, backups, replication, index maintenance on large vector tables; these are all things you end up thinking about when you should be thinking about your application.
Rivestack provides managed PostgreSQL with pgvector pre-enabled. NVMe-backed storage means your HNSW index fits in memory or loads quickly from disk, which matters for query latency. Horizontal read replicas let you offload vector search from your write path as traffic grows.
The connection string drops straight into the code above. No configuration changes needed.
Start with a Rivestack free trial and run this pipeline against a real managed database in about five minutes.
Summary
A RAG pipeline built directly on pgvector and the OpenAI API has a few real advantages over framework-based approaches:
- No abstraction layer to debug when retrieval quality degrades
- Full SQL for metadata filtering, joins, and access control
- Standard PostgreSQL tooling for backups, monitoring, and scaling
- One fewer service to operate compared to a dedicated vector store
The code in this guide is production-usable with small additions (error handling, retry logic on API calls, async ingestion). Start with this structure, measure retrieval quality against your actual questions, and add complexity only where the benchmarks tell you to.
The data you retrieve determines the answer quality far more than the model you use. Time spent improving chunking, adding metadata filtering, and tuning the HNSW index parameters is almost always better spent than switching to a larger LLM.